#' Add two numbers together.
#'
#' This function takes two numeric inputs and returns their sum.
#'
#' @param x A numeric value, the first number.
#' @param y A numeric value, the second number.
#' @return The sum of `x` and `y`.
#' @examples
#' add_numbers(5, 3)
#' add_numbers(10, -2)
add_numbers <- function(x, y) {
x + y
}Time to Roclet! admiral’s use of custom roclets to enhance and extend examples in function documentation
ABSTRACT
R package function documentation is singular in open source. Detailed argument definitions coupled with inline examples can help breakdown complicated functions that help solve problems. In admiral, R package for ADaMs, we have always prided ourselves on top-notch function documentation. However, as the examples grew for each function the documentation became unwieldy. Using the roxygen2 package, a key component of any R package, we were able to extend the roclets to better display and curate the examples for each function. A roclet is a specific parser used by the roxygen2 package in R to extract documentation from special comments within R code, automatically generating .Rd help files and other documentation components for an R package. This presentation/paper will discuss the technical details of how we extended the roclet and how this positively impacted our function examples. Improved function documentation is not only helpful to users, but also for AIs/LLMs building knowledge off your documentation.

INTRODUCTION
To set the stage, we will need to first discuss what is an R package and some of the infrastructure behind it. We will then pivot to discuss R packages that help with the package development process, i.e. roxygen2, devtools, usethis and then briefly discuss admiral. This is not a comprehensive overview. Readers are encouraged to review References for additional R package documentation.
R Packages
R packages are fundamental extensions to the R statistical programming language, bundling together code, data, documentation, and tests in a standardized format. They allow users to share and reuse code easily, greatly expanding R’s functionality and driving its widespread adoption in data science. Creating an R package typically involves using development packages like devtools and usethis to set up the necessary directory structure and compile the package. You then add your R functions to the R/ directory, write documentation for them using roxygen-style comments, and declare any package dependencies in the DESCRIPTION file. (Reference 3) After iterating through the process of generating documentation, testing your package, fixing issues and building and installing it for use, you will submit it to a package repository like CRAN. In my opinion, packages that take time to properly document their functions and build out helpful examples tend to be very successful with their user base.
roxygen2 package
I strongly encourage you to check out the R packages devtools and usethis. These two packages are incredibly helpful for the package development process. However, we will just focus on a specific package called royxgen2 for building documentation in R packages.
The roxygen2 R package simplifies the process of documenting R code within packages. (Reference 3) It allows developers to write documentation directly alongside their function definitions using special comments, known as roxygen comments, which start with #‘. Note that the package is called roxygen2 while we refer to the process of adding #’ as just roxygen or roxygenize. This approach keeps code and documentation co-located, making updates more efficient. The package roxygen2, has several functions that help process these “roxygen comments” to automatically generate .Rd files, which are the standard R documentation files. This automated generation eliminates the need to manually write complex .Rd files, significantly streamlining package development allowing developers to stay focused on writing code that helps solve a user’s problems. Beyond .Rd files, roxygen2 also helps to manage the NAMESPACE file and other package metadata, simplifying the overall package development workflow.
Below we have a brief example of a function add_numbers() with its roxygen documentation. Please note the use of #’ at each line before the function and special tags that start with @ symbol.
Let’s take a quick look at each of roxygen sections/tags:
- #’ Add two numbers together - serves as the title for the function.
- #’ This function takes two numeric inputs and returns their sum - serves as the description of the function.
- @param x A numeric value, the first number - describes the first argument, x.
- @param y A numeric value, the second number - describes the second argument, y.
- @return The sum of x and y. - describes what the function returns.
- @examples provides executable R code demonstrating how to use the function.
Typically, this function would be located in a file called add_numbers.R within a folder called /R. When a developer runs the function roxygenize() from the roxygen2 package it will create a corresponding .Rd file in /man folder. Below you will find the add_numbers.Rd. Not something you want to edit by hand!
% Generated by roxygen2: do not edit by hand
% Please edit documentation in R/add_numbers.R
\name{add_numbers}
\alias{add_numbers}
\title{Add two numbers together.}
\usage{
add_numbers(x, y)
}
\arguments{
\item{x}{A numeric value, the first number.
\describe{
\item{Default value}{none}
}}
\item{y}{A numeric value, the second number.
\describe{
\item{Default value}{none}
}}
}
\value{
The sum of \code{x} and \code{y}.
}
\description{
This function takes two numeric inputs and returns their sum.
}
\examples{
add_numbers(5, 3)
add_numbers(10, -2)
}The focus of this paper will be on enhancing the abilities of the @examples tag within the function documentation.
admiral package
The admiral R package is an open-source, modularized toolbox specifically designed for creating Analysis Data Model (ADaM) datasets within the R environment. (Reference 4) These ADaM datasets are a mandatory component for regulatory submissions to health authorities like the FDA. admiral provides a collection of reusable R functions that enable the programming community to develop CDISC-compliant ADaM datasets in a standardized and efficient manner. It promotes a “building block” approach, allowing users to combine modular functions for deriving new variables and parameters, thereby minimizing redundancy and enhancing reproducibility. The package is part of the broader pharmaverse initiative, fostering collaboration among pharmaceutical companies and streamlining the use of R for clinical trial data programming. (Reference 6)
admiral has a lot of functions to do many different derivations for ADaMs. Many functions need to be flexible to account for different scenarios, filtering or dirty data. As there are many functions with a lot of flexibility there will also be many examples. Finding a way to organize these functions in a way that helps a user is paramount to admiral’s mission. The advent of AI/LLMs that can easily harvest information makes this even more paramount, i.e. improving function documentation. Chat bots and coding agents harvesting poor documentation examples could impact the use of a package. As admiral wants to stay relevant with the times, improving our documentation for both users and AI/LLMs became a very important goal for us in 2024/2025.
R PACKAGE DOCUMENTATION
A lot of R packages have amazing documentation with great examples. However, as a package grows in the number of functions the documentation will also grow. If your package’s functions are developed in a way that the functions are flexible, then you will need to showcase this flexibility with examples. Therefore, flexibility introduces the need for more examples. To first introduce this problem with a simple example, we will look at the dplyr R package looking at the arrange() function.
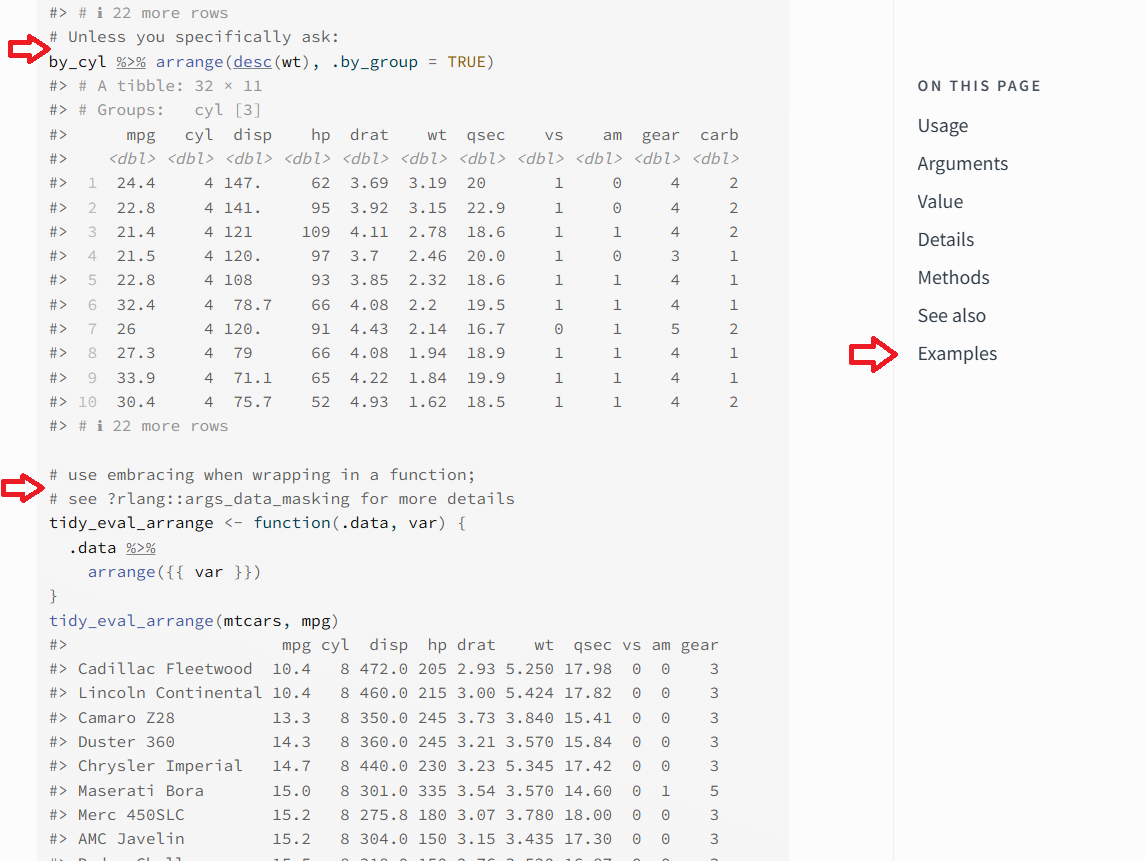
The arrange() function has limited arguments, but is very versatile and has several long examples. In the screenshot above for the dplyr arrange() documentation, the two red arrows on the left are to highlight where the use of the function changes for the examples. Code comments, #, are used to highlight what the change is and serve as a marker for a new example for the function. The red arrow on the right highlights how to get access to the examples. When you click the Example link on right-side it will take you to the top of the examples.
ADMIRAL DOCUMENTATION
The admiral R package followed a similar pattern as dplyr before the release of 1.3.0. In the next section, we will look at the same function derive_vars_dt() for how the examples changed from 1.2.0 to 1.3.0. The derive_vars_dt() function derives a date from a date character vector with the added abilities to do imputation on a partial date.
derive_vars_dt() at 1.2.0
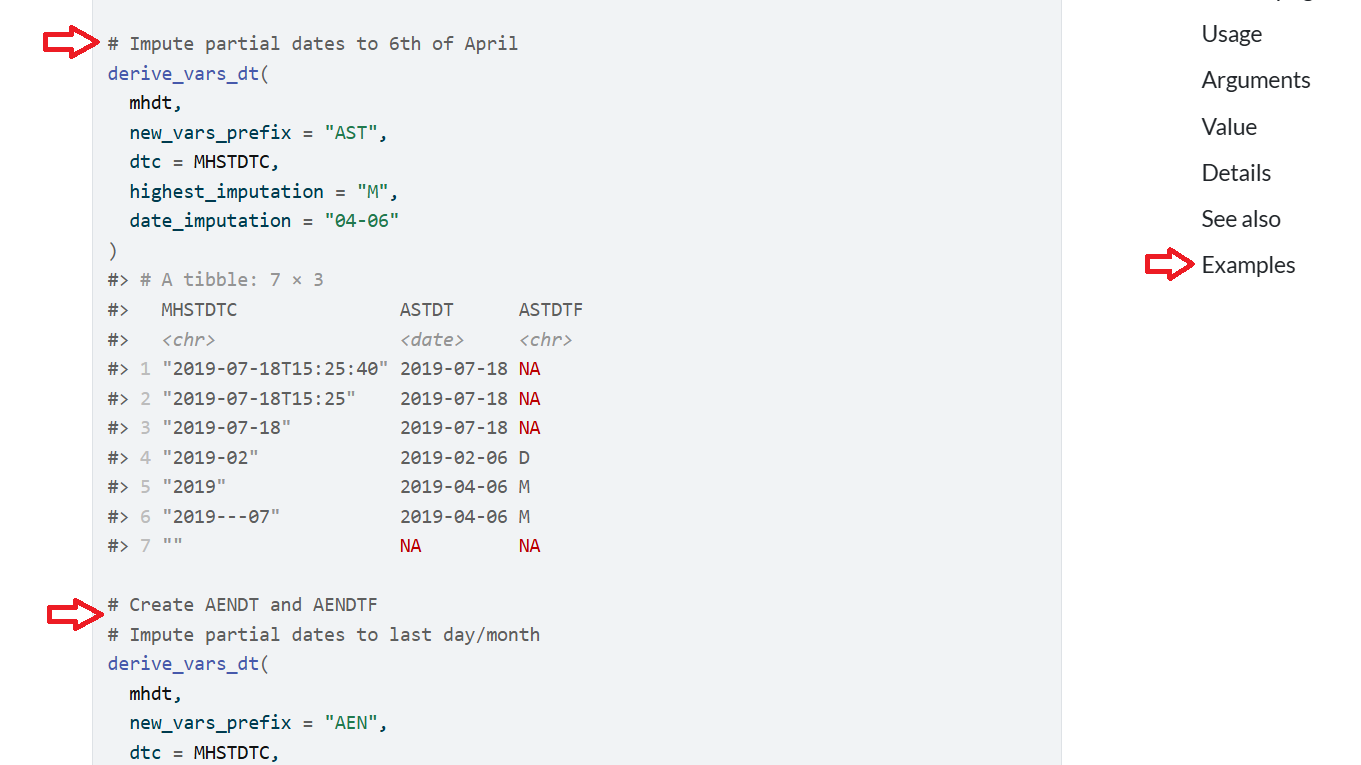
The pattern of red arrows is similar to the dplyr arrange() function. The left-side two red arrows have comments inserted to break up the different examples and show how the function can be used to do different imputations. The right-side red arrow has the link for the examples. When you click the Examples link it will take you to the top of the examples. The documentation is extensive for this function, but the display of examples is jumbled together. There is room for improvement!
derive_vars_dt() at 1.3.0
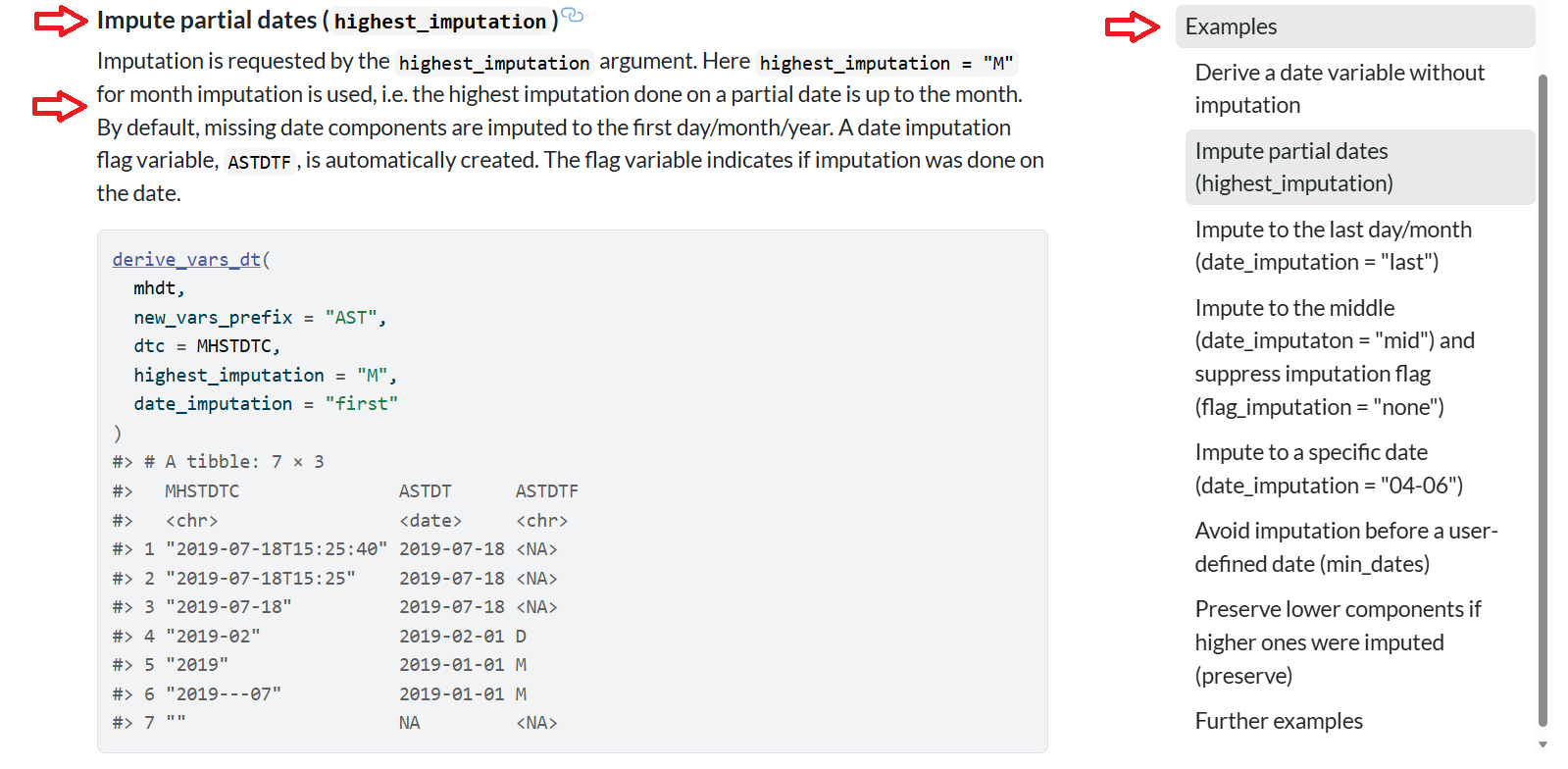
Now we are cooking! The example for derive_vars_dt() has a title for it followed by a detailed description as evidenced by the red arrows on the left side. On the right side, we can see a “Table of Contents” outlined for each example. I find this particular helpful for users as we can scaffold the examples in complexity. The examples are also clickable on the right side - so you can go to any example by just a click!
Let’s dive into how we did this, by first exploring the new roxygen tags and then get more into the code that generates the new roxygen examples.
admiral on roxygen steroids
Unfortunately, showcasing the derive_vars_dt() new roxygen layout would take up a few pages so we will use a reduced version from the admiral developer documentation, housed in a package called admiraldev, to illustrate its enhanced abilities with a function called demo_fun().
#' A Demo Function
#'
#' This function is used to demonstrate the custom tags of the `rdx_roclet()`.
#'
#' @param x An argument
#' @param number A number
#' @permitted A number
#' @param letter A letter
#' @permitted [char_scalar]
#' @default The first letter of the alphabet
#' @keywords internal
#' @examplesx
#' @info This is the introduction.
#' @caption A simple example
#' @info This is a simple example showing the default behaviour.
#' @code demo_fun(1)
#' @caption An example with a different letter
#' @info This example shows that the `letter` argument does not
#' affect the output.
#' @code demo_fun(1, letter = "b")
demo_fun <- function(x, number = 1, letter = "a") 42First, notice the use of a new roxygen tag @examplesx, which extends the abilities of the original @examples. (Reference 2) Once @examplesx is declared, you can now use the following tags:
- @caption
- @info
- @code
@caption allows for clearer titles to be declared as seen in derive_vars_dt() 1.3.0. @info allows for more detailed descriptions per example. Finally, @code is the specific code needed for the example. You can repeat this pattern to showcase many examples, like we saw for derive_vars_dt() 1.3.0 on the right-hand side of the above image. Please note that not every function in admiral has this enhancement, just ones that have many examples to showcase the range of the function.
Below you will find the .Rd portion of our demo_fun() function. This .Rd is generated in the same process as discussed above. I have removed the section pertaining to the previously discussed arguments to save some space and only showcase the sections generated for @caption, @info, @code. Again, you don’t want to manually create this either!
% Generated by roxygen2: do not edit by hand
% Please edit documentation in R/demo_fun.R
\name{demo_fun}
\alias{demo_fun}
\title{A Demo Function}
\usage{
demo_fun(x, number = 1, letter = "a")
}
...
\keyword{internal}
\section{Examples}{
This is the introduction.
\subsection{A simple example}{
This is a simple example showing the default behaviour.
\if{html}{\out{<div class="sourceCode r">}}\preformatted{demo_fun(1)
#> [1] 42}\if{html}{\out{</div>}}}
\subsection{An example with a different letter}{
This example shows that the \code{letter} argument does not affect the output.
\if{html}{\out{<div class="sourceCode r">}}\preformatted{demo_fun(1, letter = "b")
#> [1] 42}\if{html}{\out{</div>}}}}admiral roxygen behind the scenes
The admiral R package has a dependency package called admiraldev, which houses utility functions not needed by the user to do ADaM derivations. A lot of these utility functions are to check user inputs and provide feedback to the users on issues with their data. (Reference 5) The package is also home to developer utilities and so houses the roclet extension functions contained within a file called rdx_roclet.R. Readers are encouraged to review the file on the admiraldev GitHub linked in the References below. (Reference 1) The rdx_roclet.R file contains several powerful functions to extend the standard roclet, meaning the functions first performs its own custom processing first, and then delegates to the standard rd roclet to finish generating the .Rd file. This allows it to introduce new Roxygen tags and modify how existing information is presented. The roxygen2 package has a great article on extending the roclet which is recommended reading. (Reference 2)
Let’s try and break down how the rdx_roclet.R related functions work to produce the extended roxygen abilities. Note this is a high-level overview and readers are encouraged to explore the file for more details found in the References.
@examplesx (marks the beginning of the Examples Section)
Purpose: To explicitly mark the beginning of the examples section. It’s largely a structural tag and can often be omitted if the first example-related tag is @caption.
Processing: Once this @examplesx is declared, the transform_examplesx() function will look for and processes custom Roxygen tags like @caption, @info, and @code to generate structured examples. It groups these tags, executes the R code provided by @code (including capturing its output and handling expected conditions), and then combines this information into a single examplex tag for the documentation, effectively transforming descriptive and executable components into a cohesive example. Crucially, it also cleanses the executed code’s output of problematic Unicode characters to prevent LaTeX compilation errors in the final PDF manual.
@caption (Example Title)
Purpose: To provide a descriptive title or caption for a specific example within the examples section.
How it works: Each @caption begins a new example block. The text following @caption becomes the title.
Processing: transform_examplesx() collects the caption. In the final .Rd output, this becomes a \subsection{} within the main \section{Examples}{}.
@info (Example Description)
Purpose: To provide additional descriptive text or context for an example, explaining what it demonstrates or any specific considerations.
How it works: Placed after @caption and before @code, it contains the explanatory text.
Processing: transform_examplesx() appends this text to the contents of the current example block.
Note: You can add additional @info tags per example if desired. Check out the documentation for admiral derive_vars_joined().
@code (Executable Example Code)
Purpose: To provide the actual R code for an example. Critically, this code is executed during the R package check process, and its output is included in the documentation.
How it works: The R code follows the @code tag. It can also include an optional argument [expected_cnds = “warning”] to indicate that certain conditions (like warnings or errors) are anticipated from the code and should not cause the check to fail.
Processing: transform_examplesx() extracts the code and any expected_cnds options. It then calls the execute_example() function. execute_example() runs the R code in a controlled environment, captures its output (what R would print to the console), and any messages, warnings, or errors.
Note: You can add additional @code tags per example if desired. Check out the documentation for admiral derive_vars_joined().
If an unexpected condition occurs (i.e., a warning or error not listed in expected_cnds), it will throw an error, causing the R CMD check to fail. A failing R CMD check will not pass CRAN checks. This ensures that examples are not only documented but also correct and functional.
The captured code and its output are then formatted and included as part of the example’s contents.
Viola! We get pretty examples with amazing layout like below.

CONCLUSIONS
The admiral R package, which specializes in creating Analysis Data Model (ADaM) datasets, encountered a challenge with increasingly unwieldy function documentation as its examples grew in number and complexity. Coupled with the rise of AI/LLMs that harvest documentation, the admiral team looked at home to improve their documentation. To address this, admiral significantly enhanced its documentation by extending roxygen2’s roclets—a core component of R package documentation. The key innovation involved introducing custom roxygen tags: @examplesx, @caption, @info, and @code. This roxygen extension allows for a structured and user-friendly presentation of examples, as seen in the derive_vars_dt() function. Ultimately, this method enables admiral to provide scaffolding for its examples, allowing users to easily navigate and understand complex function usage, thereby significantly improving the overall user experience and the quality of its function documentation and making it easier for AIs/LLMs to harvest information.
REFERENCES
admiraldev rdx_roclet.R https://github.com/pharmaverse/admiraldev/blob/main/R/roclet_rdx.R
Extending Roclets. https://roxygen2.r-lib.org/articles/extending.html.
In-line documentation for R. (n.d.). https://roxygen2.r-lib.org/
ADAM in R Asset Library. (n.d.). https://pharmaverse.github.io/admiral
Utility functions and development tools for the Admiral Package Family. (n.d.). https://pharmaverse.github.io/admiraldev
pharmaverse (n.d.). https://pharmaverse.org/
Derive/Impute a Date from a Date Character Vector (n.d.). https://pharmaverse.github.io/admiral/reference/derive_vars_dt.html
ACKNOWLEDGMENTS
The R open-source community. You all are a special bunch.
CONTACT INFORMATION
Your comments and questions are valued and encouraged. Contact the authors at:
Author Name: Edoardo Mancini
Company: Roche
Email: edoardo.mancini_at_roche.com
Author Name: Ben Straub
Company: GSK
Email: ben.x.straub_at_gsk.com
Website: https://www.linkedin.com/in/ben-straub/
Brand and product names are trademarks of their respective companies. R